□ DGIST (President Kunwoo Lee) announced that a research team led by Prof. Yoo Wookyung (Department of Brain Sciences) and Prof. Kim Jin Hae (Department of New Biology) developed an innovative analytical technology in collaboration with a research team led by Dr. Lee Young-ho from the Protein Structure/Interaction Research Group at the Korea Basic Science Institute (KBSI). This technology enables precise, atomic-level examination of intrinsically disordered proteins (IDP), which are difficult to analyze because of their lack of fixed forms.
□ While proteins are known to perform normal functions when they possess a well-defined three-dimensional structure, approximately one-third of human proteins are IDP, which change constantly without a specific structure or shape, much like a limp skein. These proteins play essential roles, such as delivering intracellular signals, but if they are abnormally misfolded or aggregated, they become a key cause of neurodegenerative diseases (e.g., Alzheimer’s and Parkinson’s diseases, which are represented by dementia) and metabolic disorders (e.g., type 2 diabetes). However, because of the ever-changing nature of their movements, a technical limitation has existed in elucidating the specific misfolding mechanisms involved in the development of various diseases.
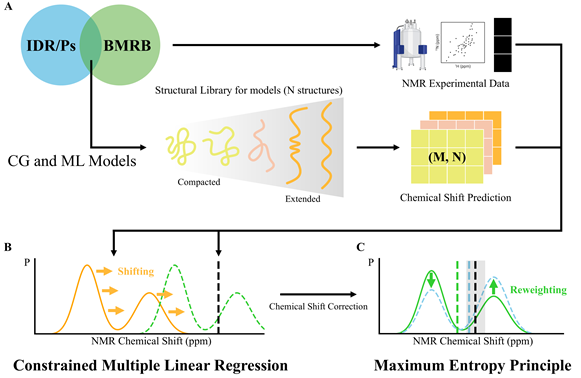
□ To overcome the limitation, the research team established an optimized fusion strategy that combined computer simulation technology with actual experimental data. First, the team generated tens of thousands of structural candidates of proteins using an artificial intelligence (AI) model, advanced simulations, and structure-related information from the Protein Data Bank (PDB).
□ Next, the research team applied a “maximum entropy” technique to compare the candidates against actual nuclear magnetic resonance spectroscopy (NMR) data obtained from real experiments and assign higher weights to structures that are closest to the actual protein state. This technology’s key feature is its ability to accurately identify even intermediate-level structures that proteins form for brief moments.
□ Notably, this achievement was made possible by the precise NMR experimental data provided by Dr. Lee Young-Ho’s research team at the KBSI. KBSI’s precise NMR data facilitated the observation of protein structures at the atomic level in the solution state rather than in the solid or crystalline state, and played a decisive role in validating and providing feedback on whether the algorithm developed by the DGIST’s research team accurately reflected the actual physical properties of proteins. Based on this, the research team tracked precisely how protein structures change depending on temperature or genetic mutations.
□ Prof. Yoo of DGIST stated, “This research is a valuable achievement resulting from the integration of DGIST’s Supercomputing AI Education and Research Center’s computational resources and advanced computational science technologies with KBSI’s world-class precision analysis infrastructure and techniques.” He added, “By unraveling the structural secrets of amorphous proteins, which were previously impossible to analyze, we expect this to become a crucial analytical tool for understanding the pathogeneses of intractable diseases, such as dementia, and for developing treatments to control them.”
□ Dr. Lee of KBSI said, “Through ongoing collaboration with DGIST, we plan to continue developing structural research tools targeting amorphous and disease-related proteins.” He added, “Together with the Korea Institute of Science and Technology Information (KISTI), we will establish a Korean version of PDB (PDBj in Korea) based on PDBj, one of the world’s top three PDBs, to create an archive for the structures of amorphous proteins lacking fixed three-dimensional structures.”
□ This research featured Jeon Joo-hyung, an integrated master’s and Ph.D. candidate at DGIST, as the first author and listed DGIST professors Yoo Wookyung and Kim Jin Hae, as well as KBSI’s Dr. Lee Young-ho, as co-corresponding authors. The research findings were published in the Proceedings of the National Academy of Sciences (PNAS), a globally authoritative science journal, on February 18. The work was supported by the National Research Foundation of Korea (NRF) and the Korea Basic Science Institute (KBSI).